This vignette reproduces the examples presented in Loureiro et al. (2026). The real-life examples
included here are the Cars and Spotify Tracks datasets, which are
discussed in Sections 6.1 and 6.2 of the paper, respectively. The
datasets are available in the package and can be loaded using
data("intCars") and
data("spotify_tracks").
The intData class is used to create objects that
represent interval-valued data. The IMCD function computes
the reweighted IMCD estimates, which are robust estimates of location
and scatter for interval-valued data. The function allows for different
cutoff methods and levels for a one-step reweighting process. The
function int_outliers identifies potential outliers in
interval-valued data using robust distance-based methods with
customizable cutoff criteria. The examples provided here demonstrate how
to apply these methods to real datasets, and the results can be compared
to those presented in the paper for validation.
For more details and examples on the intData class,
please see the vignette: Class
intData examples.
Cars Dataset
This dataset contains interval data of car specifications (Duarte Silva and Brito (2025)), including min-max values. The aggregation of the microdata was done by car model, resulting in observations. It is composed of variables:
- Engine Capacity (EngCap)
- Top Speed
- Acceleration
- Price (lnPrice after log transformation)
- Class (values: Berlina, Luxury, Sportive, and Utilitarian)
As no microdata are available, and following the standard assumption
in the literature, we adopt a continuous uniform distribution for the
latent variables, corresponding to the symmetric and i.d. case with
.
This is the default setting for the intData class.
data(intCars)
cars_microdata <- intCars$microdata
cars_int <- intCars$intDataThe IMCD function is used to compute the reweighted IMCD
estimates and the robust squared Interval-Mahalanobis distances of each
observation from the estimated barycenter. The subset size is set to
,
and the reweighting cutoff to “farness” with a cutoff level of
.
The int_outliers function identifies potential outliers
based on the robust distances obtained from the IMCD estimates, using
the same cutoff criteria.
cars_IMCD <- IMCD(cars_int, m = floor(0.75*cars_int@NObs), cutoff = "farness", cutoff_lvl = 0.9)
cars_outliers <- int_outliers(cars_IMCD$robust_dist, cutoff = "farness", cutoff_lvl = 0.9)
cars_outliers$outliers_names
#> [1] "Ferrari" "HondaNSK" "MercedesSL" "MercedesClasseS"
#> [5] "Porsche"Pairs plot, the lower triangular shows scatter plots of the four variables, with the outlying observations highlighted in red, while the upper triangular shows the interval correlation matrix.
cars_outliers_colors <- rep('gray50', cars_int@NObs)
names(cars_outliers_colors) <- rownames(cars_int)
cars_outliers_colors[cars_outliers$outliers_names] <- 'red'
SYMB.pairs.panels(cars_int, palette = cars_outliers_colors, type = "rectangles",
corr = cov2cor(cars_IMCD$cov_IMCD), labels = colnames(cars_int),
is_outlier = cars_outliers$is_outlier, gap = 0)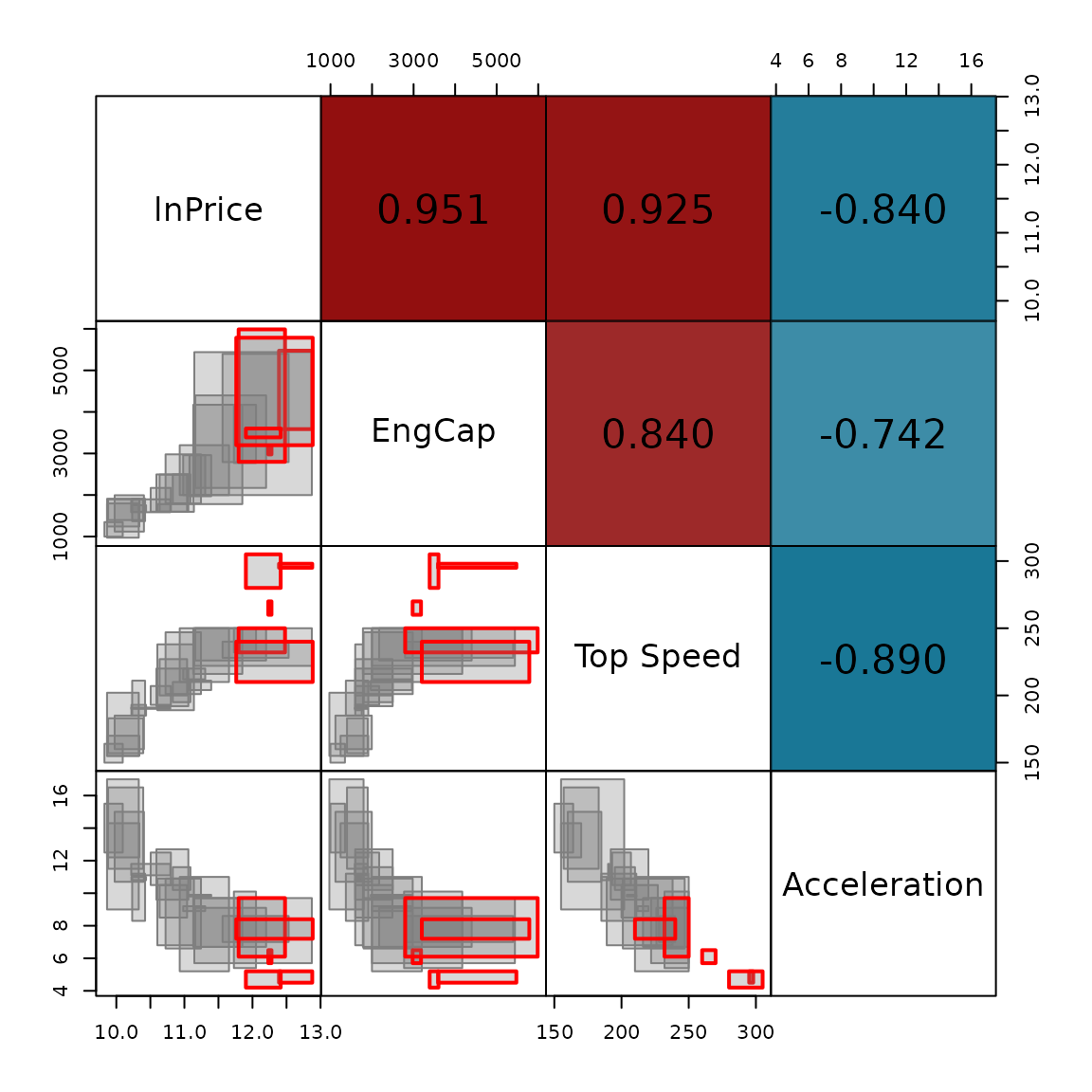
Distance-distance plot of the classical versus robust squared Interval-Mahalanobis distances. The points’ shape represents the car model class, the horizontal and vertical lines correspond to the 0.9 farness cutoff value and the 1.5 adjusted boxplot cutoff value, respectively, with the outliers marked in red.
# Classical distances and outliers
cars_class_dist <- IMah_dist(cars_int, z = rep(1,cars_int@NObs))
cars_class_outliers <- int_outliers(cars_class_dist, cutoff = "adjbox", cutoff_lvl = 1.5)
cars_is_outliers <- as.character(cars_outliers$is_outlier)
cars_is_outliers[cars_outliers$is_outlier] <- "Outlier"
cars_is_outliers[!cars_outliers$is_outlier] <- "Inlier"
plot_dist_dist(cars_class_dist, cars_class_outliers$cutoff_value[[2]], class_cutoff_label = "1.5 Adjbox",
cars_IMCD$robust_dist, cars_outliers$cutoff_value, rob_cutoff_label = "0.9 Farness",
color_class = cars_is_outliers, ggplotly = FALSE, shape_class = cars_microdata$class,
shape_label = "Class", palette = c("gray50","red"),
label_obs = c(cars_outliers$outliers_names, "Bmwserie7"))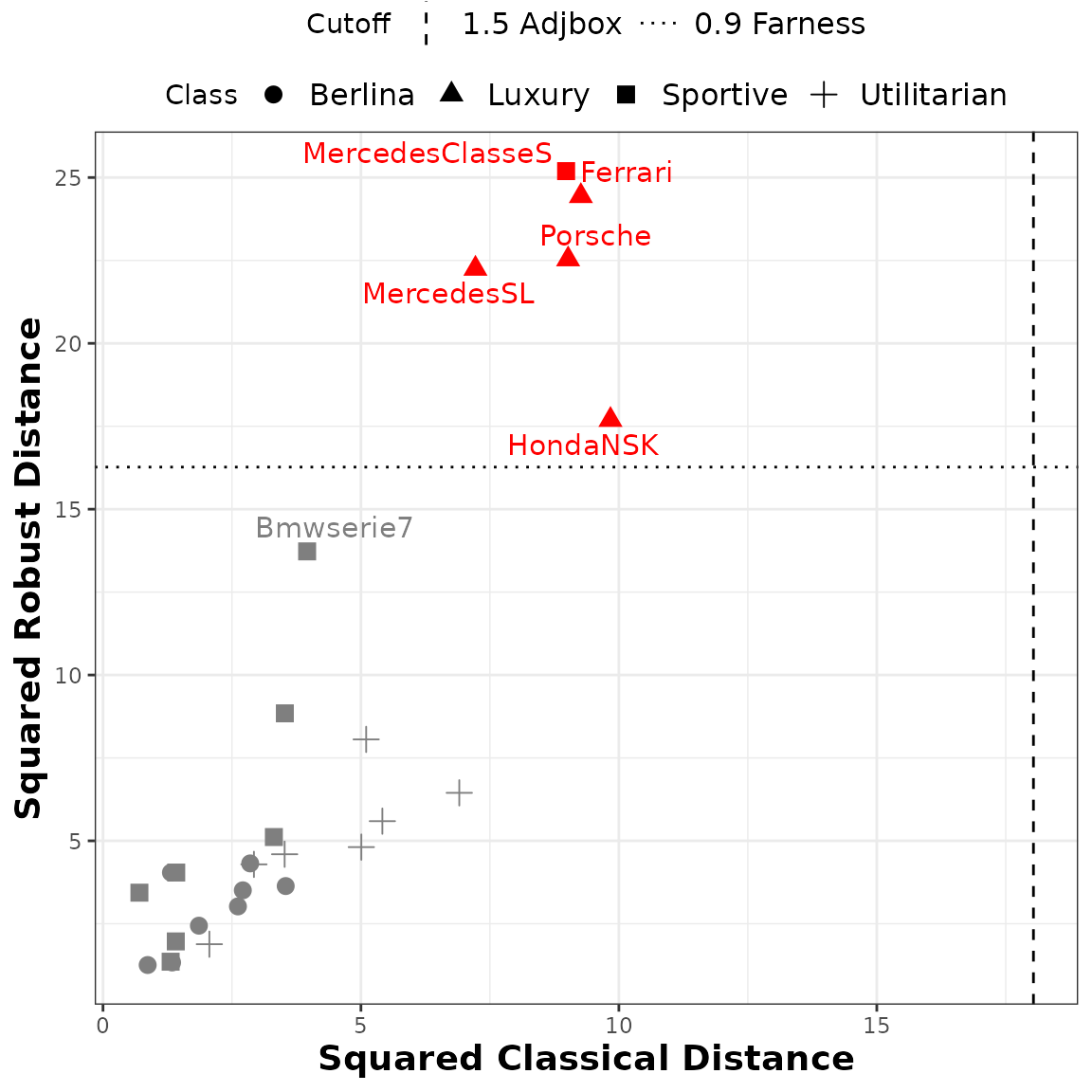
Spotify Tracks Dataset
This dataset contains interval data of Spotify tracks’ audio features (Pandya (2022)), including min-max values and trimmed intervals, as well as the microdata. The aggregation of the microdata was done by track genre, resulting in observations. It is composed of 11 audio features:
- duration
- danceability
- energy
- loudness
- speechiness
- acousticness
- instrumentalness
- liveness
- valence
- tempo
- popularity
Prior to aggregation, logarithmic transformations were applied to
loudness and tempo, duration_ms in
milliseconds was converted to duration in minutes, and
popularity was scaled to the range
.
The latent variables’ parameters are estimated from the microdata, using
Kernel Density Estimation (KDE) (package kde1d). Here, we
will use the data aggregated into trimmed intervals, taking as lower and
upper bounds the
and
quantiles, respectively.
data(spotify_tracks)
spotify_int <- spotify_tracks$intData_trimmedThe IMCD estimates are computed using the IMCD function
with a subset size of
,
and a reweighting cutoff based on “farness” with a cutoff level of
.
The int_outliers function applies the outlier detection
rule using a “farness” cutoff of
to identify strong outliers and
for mild outliers.
spotify_IMCD <- IMCD(spotify_int, m = round(0.75*nrow(spotify_int)),
cutoff = "farness", cutoff_lvl = 0.95)
# Strong outliers
spotify_outliers <- int_outliers(spotify_IMCD$robust_dist, cutoff = "farness", cutoff_lvl = 0.95)
spotify_outliers$outliers_names
#> [1] "classical" "comedy" "grindcore" "sleep"
# Mild outliers
spotify_outliers_2 <- int_outliers(spotify_IMCD$robust_dist, cutoff = "farness", cutoff_lvl = 0.9)
spotify_outliers_2$outliers_names[!spotify_outliers_2$outliers_names%in%spotify_outliers$outliers_names]
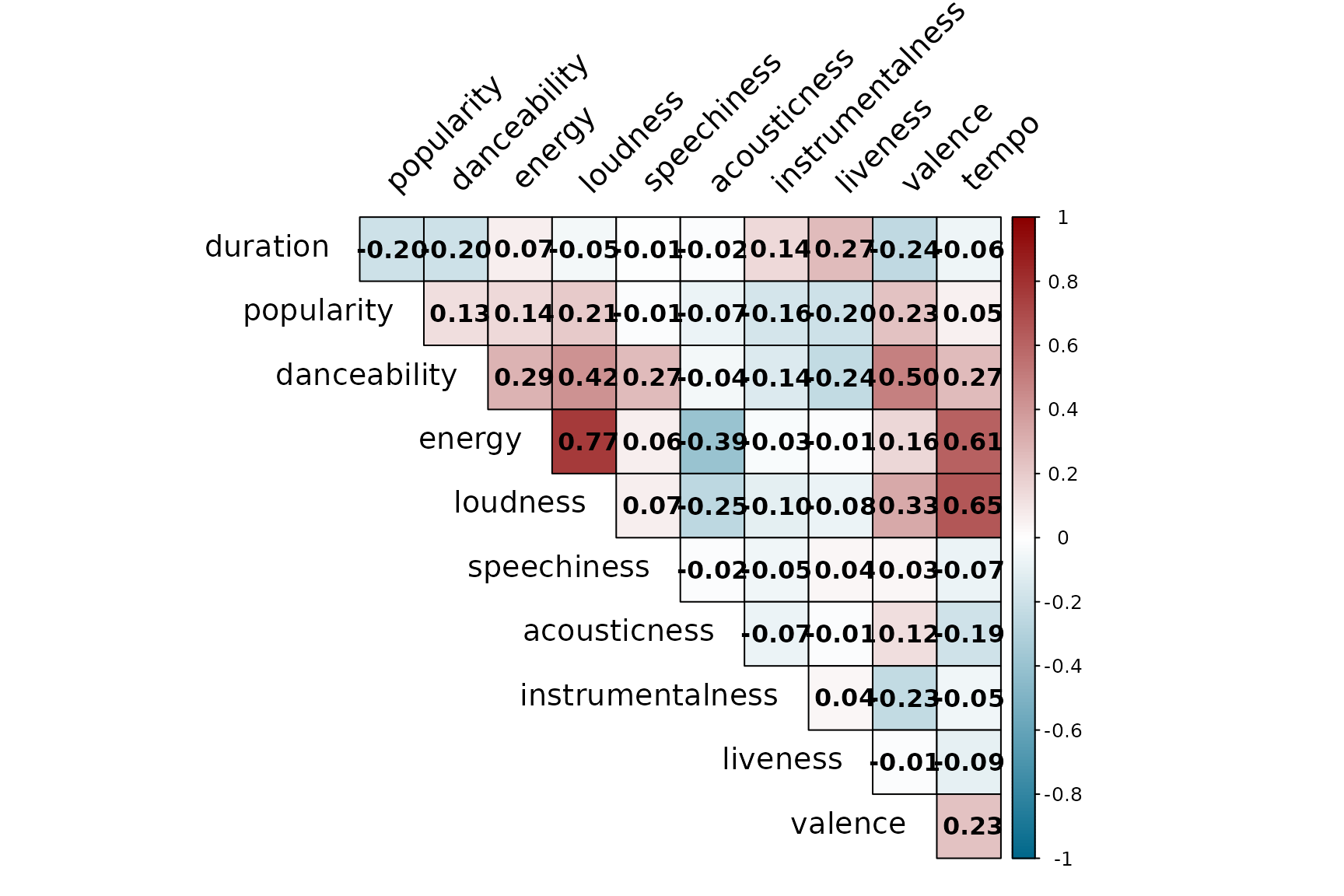
#> [1] "ambient" "chicago-house" "death-metal" "iranian"Interval correlation matrix plot based on the IMCD estimates.
# Compute correlation matrix from the robust covariance matrix
spotify_corr <- cov2cor(spotify_IMCD$cov_IMCD)
colfunc <- colorRampPalette(c("deepskyblue4", "white", "red4"))
corrplot::corrplot(
spotify_corr,
method = "color",
type = "upper",
diag = FALSE,
col = colfunc(200),
tl.col = "black",
tl.srt = 45,
tl.offset = 1,
tl.cex = 1.2,
outline = TRUE,
addCoef.col = "black",
number.cex = 1
)
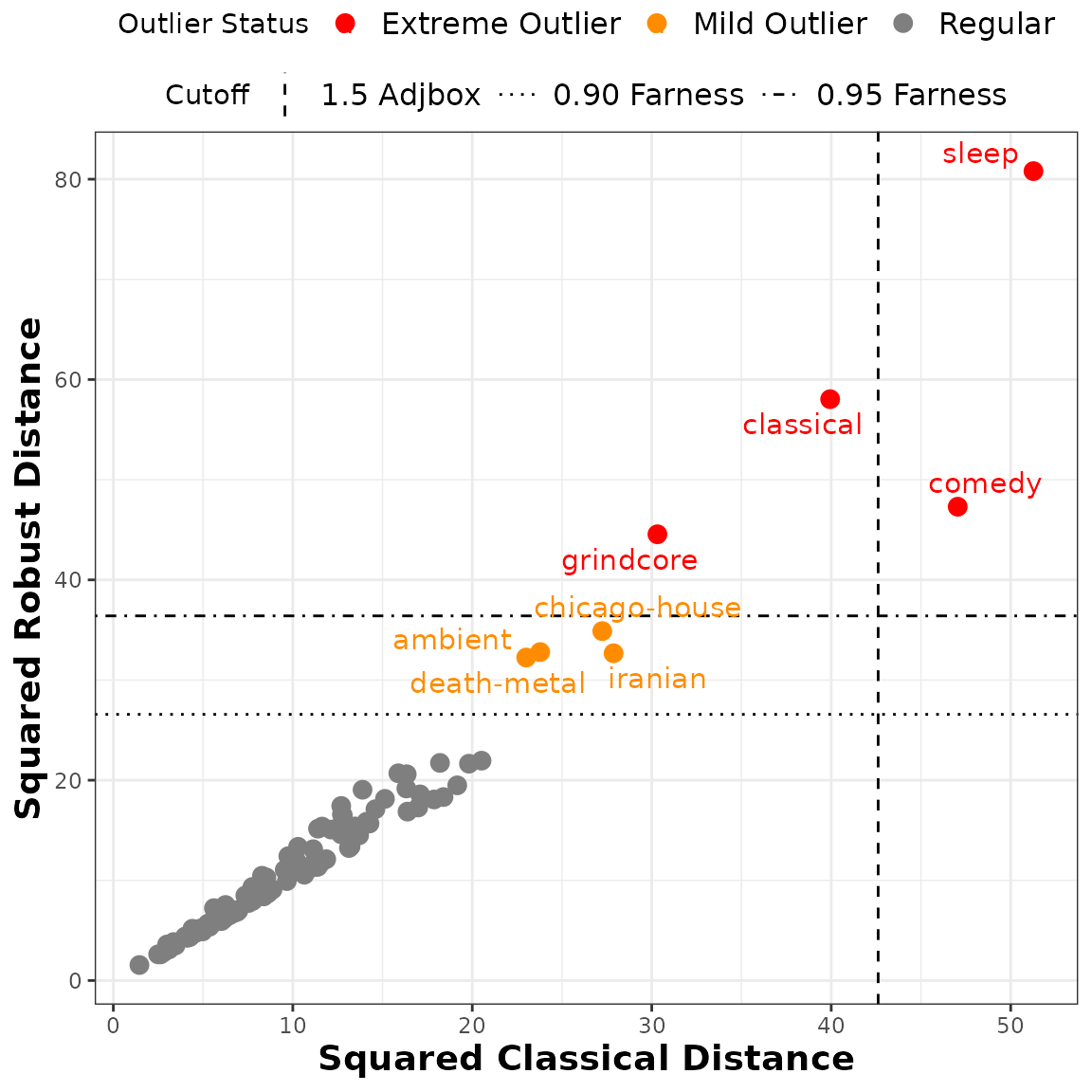
Distance-distance plot of the classical versus robust squared Interval-Mahalanobis distances for the Spotify dataset. The horizontal lines correspond to the and farness cutoff values, with the extreme outliers marked in red and the mild outliers in orange, while the vertical line represents the adjusted boxplot cutoff.
# Classical distances and outliers
spotify_class_dist <- IMah_dist(spotify_int, z = rep(1,spotify_int@NObs))
spotify_class_outliers <- int_outliers(spotify_class_dist, cutoff = "adjbox")
spotify_is_outliers <- as.character(spotify_outliers$is_outlier)
spotify_is_outliers[!spotify_outliers_2$is_outlier] <- "Regular"
spotify_is_outliers[spotify_outliers_2$is_outlier] <- "Mild Outlier"
spotify_is_outliers[spotify_outliers$is_outlier] <- "Extreme Outlier"
palette_outliers <- c(
"Regular" = "gray50",
"Mild Outlier" = "darkorange",
"Extreme Outlier" = "red"
)
plot_dist_dist(spotify_class_dist, spotify_class_outliers$cutoff_value[[2]],
"1.5 Adjbox", spotify_IMCD$robust_dist,
c(spotify_outliers_2$cutoff_value, spotify_outliers$cutoff_value),
c("0.90 Farness", "0.95 Farness"), color_class = spotify_is_outliers,
color_label = "Outlier Status", palette = palette_outliers, ggplotly = FALSE,
label_obs = spotify_outliers_2$outliers_names)